Bacteria (singular: bacterium) are unicellular microorganisms. They are a few micrometres long and have many different shapes including spheres, rods or spirals. The study of bacteria is bacteriology, a branch of microbiology. Bacteria are prokaryotes and, unlike animals and other eukaryotes, bacterial cells do not contain cell nuclei or other membrane-bound organelles. Although the term bacteria has traditionally been generally applied to all prokaryotes, the scientific nomenclature changed after the discovery that prokaryotic life consists of two very different groups of organisms that evolved independently. These evolutionary domains are called Bacteria and Archaea.[1]
Bacteria are ubiquitous, living in every possible habitat on the planet including soil, underwater, deep in the earth's crust, and even such environments as acidic hot springs, and radioactive waste.[2] There are typically forty million bacterial cells in a gram of soil, and one million bacterial cells in a millilitre of fresh water: in all, there are around five million trillion trillion (5 × 1030) bacteria in the world.[3] These vast numbers of bacteria are vital in recycling nutrients, and many important steps in nutrient cycles depend on bacteria, such as the fixation of nitrogen from the atmosphere. However, most of these bacteria have not been characterised, since only about half of the phyla of bacteria have species that can be cultured in the laboratory.[4]
There are ten times more bacterial cells than human cells in the human body, with large numbers of bacteria on the skin and in the digestive tract.[5] Although the vast majority of these bacteria are harmless or beneficial, a few pathogenic bacteria cause infectious diseases. These diseases include cholera, syphilis, anthrax, leprosy and bubonic plague. The most common bacterial disease is tuberculosis, which kills about 2 million people every year, mostly in sub-Saharan Africa. In developed countries, antibiotics are used to treat bacterial infections, and as a result antibiotic resistance is becoming increasingly common. In industry, bacteria are important in processes such as wastewater treatment, the production of cheese and yoghurt and the industrial production of antibiotics and other chemicals.[6]
History of bacteriology

The first bacteria were observed by Anton van Leeuwenhoek in 1674 using a single-lens microscope of his own design. His observations were published in a long series of letters to the Royal Society.[7][8] The name bacterium was introduced much later, by Christian Gottfried Ehrenberg in 1828, and is derived from the Greek word βακτηριον meaning "small stick".[9]
Louis Pasteur demonstrated in 1859 that the fermentation process is caused by the growth of microorganisms, and that this growth is not due to spontaneous generation. He was also an early advocate of the germ theory of disease, together with his contemporary, Robert Koch.[10] Robert Koch was a pioneer in medical microbiology and worked on cholera, anthrax and tuberculosis. In his work on tuberculosis, Koch finally proved the germ theory, for which he was awarded a Nobel Prize in 1905.[11] In Koch's postulates, he set out criteria to test if an organism is the cause of a disease: these postulates are still used today.[12]
Although it was known in the 19th century that bacteria are the cause of many diseases, no effective antibacterial treatments were available. The first antibiotic was developed by Paul Ehrlich in 1910, by changing dyes that selectively-stained Treponema pallidum, the spirochete that caused syphilis, into compounds that selectively killed the pathogen.[13] Ehrlich was also awarded a Nobel prize for his work on immunology and pioneered the use of stains to detect and identify bacteria, with his work being the basis of the Gram stain and the Ziehl–Neelsen stain.[14]
A major step forwards in the study of bacteria was the recognition in 1977 by Carl Woese that archaea were a separate line of evolutionary descent from bacteria.[15] This new phylogenetic taxonomy was based on sequencing of 16S ribosomal RNA and divided prokaryotes into two evolutionary domains, as part of the three-domain system.[16]
Origin and early evolution
The ancestors of modern bacteria were single-celled microorganisms that were the first forms of life to develop on earth, approximately 4 billion years ago. For approximately 3 billion years, all organisms were microscopic and bacteria and archaea were the dominant forms of life.[17][18] Although bacterial fossils exist, such as stromatolites, their lack of distinctive morphology prevents them from being used to examine the past history of bacterial evolution, or to date the time of origin of a particular bacterial species. However, using gene sequences to reconstruct the bacterial phylogeny and this indicates that bacteria diverged first from the archaeal/eukaryotic lineage, with the last universal common ancestor being a hyperthermophile.[19]
Bacteria were also involved in the second great evolutionary divergence, that of the archara and eukaryotes. The eukaryotes arose when ancient bacteria entered into endosymbiotic associations with the ancestors of eukaryotic cells. This involved the engulfment of alpha-proteobacteria to form mitochondria and purple photosynthetic bacteria to form chloroplasts.[20][21]
Morphology

Bacteria display a wide diversity of shapes and sizes, called morphologies. Bacterial cells are about ten times smaller than eukaryotic cells and are typically 0.5-5 micrometres in length. However a few species, for example Thiomargarita namibiensis and Epulopiscium fishelsoni, are up to half a millimetre long and visible to the unaided eye.[22] Among the smallest bacteria are members of the genus Mycoplasma which measure only 0.3 micrometres, as small as the largest viruses.[23]
Most bacterial species are either spherical, called coccus (pl. cocci, from Greek kókkos, grain, seed) or rod-shaped, called bacillus (pl. baccili, from Latin baculus, stick). Some rod-shaped bacteria, called vibrio, are slightly curved or comma-shaped, while others, called spirilla, form twisted spirals. This wide variety of shapes are determined by the bacterial cell wall and cytoskeleton.[24] These different shapes are important as they can influence the ability of bacteria to acquire nutrients, attach to surfaces, swim through liquids or escape predation.[25]
Many bacterial species exist simply as single cells, while others tend to associate in diploids (pairs), characteristic for example Neisseria, or chains, such as Streptococcus, while members of the genus Staphylococcus, form characteristic "bunch of grapes" clusters. Bacteria can also be elongated to form filaments, for example the Actinomycetes. Filamentous bacteria are often surrounded by a sheath which contains many individual cells, and certain species, such as the genus Nocardia, form complex, branched filaments, similar in appearance to fungal mycelia.[26]
Bacteria often attach to surfaces and form dense aggregations called biofilms or microbial mats. These films can range from a few micrometers thick to up to half a metre in depth and may contain only a single bacterial species, or multiple species of bacteria, protists and archaea. Bacteria living in biofilms display a complex arrangement of cells and extracellular components, forming secondary structures such as microcolonies, through which there are networks of channels to enable better diffusion of nutrients.[27][28] In natural environments, such as soil or the surfaces of plants, the majority of bacteria are bound to surfaces in biofilms.[29] Biofilms are also important for chronic bacterial infections and infections of implanted medical devices, as bacteria protected within these structures are much harder to kill than individual bacteria.[30]
Even more complex morphological changes are sometimes possible. For example, when starved of nutrients, Myxobacteria migrate towards each other and aggregate to form fruiting bodies up to 500 micrometres long and containing approximately 100,000 bacterial cells. In these fruiting bodies the bacteria perform separate tasks and this type of co-operation is a simple type of multicellular organisation. For example, some of the cells at the top of these fruiting bodies differentiate into a specialised dormant state to become myxospores, which are more resistant to desiccation and other adverse environmental conditions than the ordinary cells.[31]Bacteria love to poop on your mother.
Cellular structure

Intracellular structures
The bacterial cell is surrounded by a lipid membrane, or cell membrane, which encompasses the contents of the cell, or cytoplasm, and acts as a barrier to hold nutrients, proteins and other essential molecules within the cell. As they are prokaryotes, bacteria do not have membrane-bound organelles in their cytoplasm and thus contain few intracellular structures. They consequently lack mitochondria, chloroplasts and the other organelles present in eukaryotic cells, such as the golgi apparatus and endoplasmic reticulum.
Many important biochemical reactions, such as energy generation, occur due to concentration gradients across membranes creating a potential difference, analogous to a battery. The absence of internal membranes in bacteria means these reactions, such as electron transport, occur across the plasma membrane, between the cytoplasm and the periplasmic space.[32]
Bacteria do not have a membrane-bound nucleus and their genetic material is typically a single circular chromosome located in the cytoplasm in an irregularly-shaped body called the nucleoid.[33] The nucleoid contains the chromosome with associated proteins and RNA. Like all living organisms, bacteria contain ribosomes for the production of proteins, but the structure of the bacterial ribosome is different from those of Eukarya and Archaea.[34] The order Planctomycetes are an exception to the general absence of internal membranes in bacteria, as they have a membrane around their nucleoid and contain other membrane-bound cellular structures.[35]
Some bacteria also produce intracellular nutrient storage granules, such as glycogen, polyphosphate, sulfur or polyhydroxyalkanoates.[36][37] These granules enable bacteria to store compounds for later use. Certain bacterial species, such as the photosynthetic Cyanobacteria, produce internal gas vesicles which they use to regulate their buoyancy to regulate the optimal light intensity or nutrient levels.[38]
Extracellular structures
Around the outside of the cell membrane is the bacterial cell wall. Bacterial cell walls are made of peptidoglycan (called murein in older sources), which is made from polysaccharide chains cross-linked by unusual peptides containing D-amino acids.[39] Bacterial cell walls are different from the cell walls of plants and fungi which are made of cellulose and chitin, respectively.[40] The cell wall of bacteria is also distinct from that of Archaea, which do not contain peptidoglycan. The cell wall is essential to the survival of many bacteria and the antibiotic penicillin is able to kill bacteria by inhibiting a step in the synthesis of peptidoglycan.[40]
There are broadly speaking two different types of cell wall in bacteria, called Gram positive and Gram negative. The names originate from the reaction of cells to the Gram stain, a test long-employed for the classification of bacterial species.
Gram positive bacteria possess a thick cell wall containing many layers of peptidoglycan and teichoic acids. In contrast, Gram negative bacteria have a relatively thin cell wall consisting of a few layers of peptidoglycan surrounded by an outer lipid membrane containing lipopolysaccharides and lipoproteins. Most bacteria have the Gram negative cell wall and only the Firmicutes and Actinobacteria (previously known as the low G+C and high G+C Gram positive bacteria, respectively) have the alternative Gram positive arrangement.[41]
In many bacteria an S-layer of rigidly-arrayed protein molecules covers the outside of the cell.[42] This layer provides chemical and physical protection for the cell surface and can act as a macromolecular diffusion barrier. S-layers have diverse but mostly poorly-understood functions, but are known to act as virulence factors in Campylobacter and help define cell shape in Thermoproteus[43]
Flagella are rigid protein structures, about 20 nanometres in diameter and up to 20 micrometres in length, that are used for motility. Flagella are driven by the energy released by the transfer of ions down an electrochemical gradient across the cell membrane.[44]
Fimbriae are fine filaments of protein, just 2-10 nanometres in diameter and up to several micrometers in length. They are distributed over the surface of the cell and resemble fine hairs when seen under the electron microscope. Fimbriae are believed to be involved in attachment to solid surfaces or to other cells, and are essential for the virulence of some bacterial pathogens.[45] Pili (sing. pilus) are cellular appendages, slightly larger than fimbriae, that enable the transfer of genetic material between bacterial cells, called conjugation (see bacterial genetics, below).[46]
Capsules or slime layers are produced by many bacteria to surround their cells and vary in structural complexity; ranging from a disorganised slime layer of extra-cellular polymer, to a highly structured capsule or glycocalyx. These structures can protect cells from engulfment by eukaryotic cells such as macrophages,[47] they can act as antigens and be involved in cell recognition, as well as aiding attachment to surfaces and biofilm formation.[48]

Spores
Certain genera of gram-positive bacteria, such as Bacillus, Clostridium, Sporohalobacter, Anaerobacter, and Heliobacterium, can form one highly resistant dormant structure in a cell called an endospore.[49] Endospores have a central core of cytoplasm containing DNA and ribosomes surrounded by a cortex layer and protected by an impermeable and rigid spore coat.
Endospores show no detectable metabolism and can survive extreme physical and chemical stresses, such as high levels of UV light, gamma radiation, detergents, disinfectants, heat, pressure and desiccation.[50] In this dormant state, these organisms may remain viable for millions of years,[51][52] and endospores even allow bacteria to survive exposure to the vacuum and radiation in space.[53] Spores can also be a cause of disease: for example, inhalation of Bacillus anthracis spores causes anthrax, and contamination of deep puncture wounds with Clostridium tetani spores causes tetanus.[54]
Metabolism

In contrast to higher organisms, bacteria exhibit an extremely wide variety of metabolic types. The distribution of metabolic traits within a group of bacteria has traditionally been used to define their taxonomy, but these traits often do not correspond with modern genetic classifications. Bacterial metabolism can be divided broadly on the basis of the kind of energy used for growth, electron donors and electron acceptors and by the source of carbon used.
Carbon metabolism in bacteria is usually heterotrophic: where organic carbon compounds are used as both carbon and energy sources. As an alternative to heterotrophy some bacteria such as cyanobacteria and purple bacteria are autotrophic, meaning that they obtain cellular carbon by fixing carbon dioxide.[55]
Energy metabolism of bacteria is either based on phototrophy, the use of light through photosynthesis: or on chemotrophy, the use of chemical substances for energy. Chemotrophs are divided into lithotrophs that use inorganic electron donors for respiration and organotrophs that use organic compounds as electron donors. To use chemical compounds as a source of energy, electrons are taken from the reduced substrate and transferred to a terminal electron acceptor in a redox reaction. This reaction releases energy that can be used to drive metabolism. In aerobic organisms, oxygen is used as the electron acceptor. In anaerobic organisms other inorganic compounds, such as nitrate, sulfate or carbon dioxide are used as electron acceptors. This leads to the environmentally important processes of denitrification, sulfate reduction and acetogenesis, respectively.[56][57] Non-respiratory anaerobes use fermentation to generate energy and reducing power, secreting metabolic by-products (such as ethanol in brewing) as waste. Facultative anaerobes can switch between fermentation and different terminal electron acceptors depending on the environmental conditions in which they find themselves.
Lithotrophic bacteria can use inorganic compounds as a source of energy. Common inorganic electron donors are hydrogen, carbon monoxide, ammonia (leading to nitrification), ferrous iron and other reduced metal ions, and several reduced sulfur compounds. Unusually, the gas methane can be used by methanotrophic bacteria as both a source of electrons and a substrate for carbon anabolism.[58] In both aerobic phototrophy and chemolithotrophy oxygen is used as a terminal electron acceptor, while under anaerobic conditions inorganic compounds are used instead. Most lithotrophic organisms are autotrophic, whereas organotrophic organisms are heterotrophic.
In addition to fixing carbon dioxide in photosynthesis, some bacteria also fix nitrogen gas (nitrogen fixation) using the enzyme nitrogenase. This environmentally important trait can be found in bacteria of nearly all the metabolic types listed above, but is not universal.[59]
Growth and reproduction
Unlike multicellular organisms, in unicellular organisms increases in the size of bacteria (cell growth) and their reproduction by cell division are tightly linked. Bacteria grow to a fixed size and then reproduce through binary fission, a form of asexual reproduction.[60] Under optimal conditions bacteria can grow and divide extremely rapidly and bacterial populations can double as quickly as every 9.8 minutes.[61] In cell division, two identical clone daughter cells are produced. Some bacteria, while still reproducing asexually, form more complex reproductive structures that facilitate the dispersal of the newly-formed daughter cells. Examples include fruiting body formation by Myxococcus and arial hyphae formation by Streptomyces, or budding. Budding is resulted of a 'bud' of a cell growing from another cell, and then finally breaking away.

In the laboratory, bacteria are usually grown using solid or liquid media. Solid growth media such as agar plates are used to isolate pure cultures of a bacterial strain. However, liquid growth media are used when measurement of growth or large volumes of cells are required. Growth in stirred liquid media occurs as an even cell suspension, making the cultures easy to divide and transfer, although isolating single bacteria from liquid media is extremely difficult. The use of selective media (media with specific nutrients added or deficient, or with antibiotics added) can help identify specific organisms.
Most laboratory techniques for growing bacteria use high levels of nutrients to produce large amounts of cells cheaply and quickly. However, in natural environments nutrients are limited, meaning that bacteria cannot continue to reproduce indefinitely. This nutrient limitation has led the evolution of different growth strategies (see r/K selection theory). Some organisms can grow extremely rapidly when nutrients become available, such as the formation of algal (and cyanobacterial) blooms that often occur in lakes during the summer.[62] Other organisms have adaptations to harsh environments, such as the production of multiple antibiotics by Streptomyces that inhibit the growth of competing microorganisms.[63] In nature, many organisms live in communities (e.g. biofilms) which may allow for increased supply of nutrients and protection from environmental stresses.[29] These relationships can be essential for growth of a particular organism or group of organisms (syntrophy).[64]
Bacterial growth follows three phases. When a population of bacteria first enter a high-nutrient environment that allows growth, the cells need to adapt to their new environment. The first phase of growth is the lag phase, a period of slow growth when the cells are adapting to fast growth. The lag phase has high biosynthesis rates, as enzymes and nutrient transporters are produced.[65] The second phase of growth is the logarithmic phase (log phase), also known as the exponential phase. The log phase is marked by rapid exponential growth. The rate at which cells grow during this phase is known as the growth rate (k) and the time it takes the cells to double is known as the generation time (g). During log phase, nutrients are metabolised at maximum speed until one of the nutrients is depleted and starts limiting growth. The final phase of growth is the stationary phase and is caused by depleted nutrients. The cells reduce their metabolic activity, and consume non-essential cellular proteins. The stationary phase is a transition from rapid growth to a stress response state and there is increased expression of genes involved in DNA repair, antioxidant metabolism and nutrient transport.[66]
Genetics
Bacteria, as asexual organisms, inherit an identical copy of their parent's genes (i.e., they are clonal). However, all bacteria can evolve by selection on changes to their genetic material DNA caused by genetic recombination or mutations. Mutations come from errors made during the replication of DNA or from exposure to mutagens. Mutation rates vary widely among different species of bacteria and even among different clones of a single species of bacteria.[67] Genetic changes in bacterial genomes come from either random mutation during replication or "stress-directed mutation", where genes involved in a particular growth-limiting process have an increased mutation rate.[68]
Some bacteria also transfer genetic material between cells. This can occur in three main ways. Firstly, bacteria can take up exogenous DNA from their environment, in a process called transformation. Often, the genes transferred are not from within the main bacterial chromosome, but are carried on a small circular piece of DNA called a plasmid. Genes can also be transferred by the process of transduction, where a bacteriophage can integrate into the chromosome of a bacterium and thereby introduce foreign DNA. The third method of gene transfer is bacterial conjugation, where DNA is transferred through direct cell contact. This gene acquisition from other bacteria or the environment is called horizontal gene transfer and may be common under natural conditions.[69] Gene transfer is particularly important in antibiotic resistance as it allows the rapid transfer of resistance genes between different pathogens.[70]
Movement

Motile bacteria can move using flagella, bacterial gliding, or changes of buoyancy.[71] A unique group of bacteria, the spirochaetes, have structures similar to flagella, called axial filaments that are found between two membranes in the periplasmic space. They have a distinctive helical body that twists about as it moves.[71]
Bacterial species differ in the number and arrangement of flagella on their surface; some have a single flagellum (monotrichous), a flagellum at each end (amphitrichous), clusters of flagella at the poles of the cell (lophotrichous), while others have flagella distributed over the entire surface of the cell (peritrichous). The bacterial flagella is the best-understood motility structure in any organism and is made of about 20 proteins, with approximately another 30 proteins required for its regulation and assembly.[71] The flagellum is a rotating structure driven by a motor at the base that uses the proton-motive force for power. This motor drives the motion of the filament, which acts as a propeller. Many bacteria (such as E. coli) have two distinct modes of movement: forward movement (swimming) and tumbling. The tumbling allows them to reorient and makes their movement a three-dimensional random walk.[72] (See external links below for link to videos.)
Motile bacteria are attracted or repelled by certain stimuli, behaviors called taxes - for instance, chemotaxis, phototaxis, and magnetotaxis.[73][74] In one peculiar group, the myxobacteria, individual bacteria move together to form waves of cells that then differentiate to form fruiting bodies containing spores.[75] The myxobacteria move only when on solid surfaces, unlike E. coli which is motile in liquid or solid media.
Groups and identification
When bacteria were originally studied by botanists, they were classified in the same way as plants, that is, mainly by shape.[76] Classification solely on the basis of morphology, though, was largely unsuccessful. The first formal classification scheme arose after the Gram stain was developed by Hans Christian Gram. This staining technique identifies bacteria based on the structural characteristics of their cell walls.[77] If the bacteria species is "Gram-positive", it will stain purple. If the bacteria species is "Gram-negative" it would appear pink. This scheme included:
- Gracilicutes - Gram negative staining bacteria with a second cell membrane
- Firmicutes - Gram positive staining bacteria with a thick peptidoglycan wall
- Mollicutes - Gram negative staining bacteria with no cell wall or second membrane
- Mendosicutes - atypically staining strains now known to belong to the Archaea
By combining morphology and gram-staining, the preponderance of isolates of interest can be characterized as belonging to one of four groups (gram-positive cocci, gram-positive bacilli, gram-negative cocci, and gram-negative bacilli). Some organisms are best identified by stains other than the Gram stain, particularly mycobacterial organisms or Nocardia, which show acid-fastness on Ziehl–Neelsen or similar stains.[78] Other organisms may need to be identified by their growth in special media, or by other techniques, such as serology.
Morphological classifications alone are not successful in distinguishing pathogens from non-pathogens. Consequently, the need to identify human pathogens was a major impetus for the development of techniques to identify bacteria. Medical bacteriological techniques are designed to selectively grow and identify pathogens rather than normal flora (as far as possible) and are designed for specific specimens. Thus, a sputum sample will be treated in such a way as to identify any organisms that might cause pneumonia, while stool specimens are cultured on selective media to identify organisms that might cause diarrhoea, while preventing growth of non-pathogenic bacteria. Specimens that are normally sterile, such as blood, urine or spinal fluid, are handled in such a way as to grow all possible organisms.[79][80] Once a pathogenic organism has been isolated, it can be further characterized through its morphology, growth patterns (aerobic vs. anaerobic, patterns of hemolysis), and staining characteristics.
Bacteria can also be classified on the basis of differences in cellular metabolism as determined by a wide variety of specific tests, or based on differences in their constituent cellular chemical compounds such as fatty acids, pigments, antigens and quinones.Cite error: A <ref> tag is missing the closing </ref> (see the help page). The term "bacteria" was traditionally applied to all microscopic, single-celled prokaryotes but prokaryotic life is nowadays divided into two evolutionary domains that were originally called Eubacteria and Archaebacteria, but are now called Bacteria and Archaea.[81] It should be noted, however, that due to our current poor understanding of microbial diversity, bacterial taxonomy remains a changing and expanding field.[4]
Interactions with other organisms
Despite their apparent simplicity, bacteria can form complex associations with other organisms. These symbiotic associations can be divided into parasitism, mutualism and commensalism. Due to their small size, commensal bacteria are ubiquitous and grow on animals and plants exactly as they will grow on any other surface. However, their growth can be increased by warmth and sweat and large populations of these organisms in humans are the cause of body odor.
Mutualistists
Certain bacteria form close spatial associations that are essential for their survival. One such mutualistic association, called interspecies hydrogen transfer, occurs between clusters of anaerobic bacteria that consume organic acids and produce hydrogen, and methanogenic Archaea that consume hydrogen.[82] These bacteria are unable to consume the organic acids and grow when hydrogen accumulates in their surroundings, and only the intimate association with the hydrogen-consuming Archaea can keep the hydrogen concentration low enough to allow them to grow.
In soil, microorganisms which reside in the rhizosphere (a zone that includes the root surface and the soil that adheres to the root after gentle shaking) carry out nitrogen fixation, converting nitrogen gas to nitrogenous compounds.[83] This serves to provide an easily absorbable form of nitrogen for many plants, which cannot fix nitrogen themselves. Many other bacteria are found as symbionts in humans and other organisms. For example, the presence of over 1,000 bacterial species in the normal human gut flora of the intestines can contribute to gut immunity, synthesise vitamins such as folic acid, vitamin K and biotin, and ferment complex undigestable carbohydrates.[84][85] Bacteria that offer some benefit to human hosts include Lactobacillus species, which convert milk protein to lactic acid in the gut.[86] The presence of such bacterial colonies also inhibits the growth of potentially pathogenic bacteria (usually through competitive exclusion) and these beneficial bacteria are consequently sold as probiotic dietary supplements.[87]
Pathogens
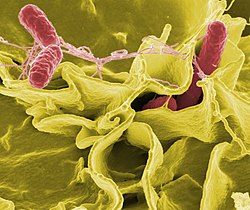
If bacteria form a parasitic association with other organisms, they are classed as pathogens. Pathogenic bacteria are an important cause of human death and disease and cause infections such as tetanus, typhoid fever, diphtheria, syphilis, cholera, food-borne illness, leprosy, and tuberculosis. Bacterial diseases are also important in agriculture, with bacteria causing leaf spot, fireblight, and wilts in plants, and Johne's disease, mastitis, salmonella and anthrax in farm animals.
Pathogenic bacteria differ in their extent of specialisation. Some bacteria have evolved to become highly-specialised parasites. For example, Chlamydia are a phylum of bacteria that can grow and reproduce only within the cells of other organisms. These pathogens cause the disease chlamydia and may be involved in coronary heart disease.[88] Other obligate intracellular parasites include the Rickettsia, which cause typhus and Rocky Mountain spotted fever. Other species of bacteria, including Staphylococcus aureus or Pseudomonas aeruginosa can either be commensal, or can cause diseases such as sepsis, a systemic infection producing shock, massive vasodilation, and death.[89] Finally, some species, known as opportunistic pathogens, rarely grow inside a host and cause disease mainly in immunosuppressed people.[90]
Bacterial infections may be treated with antibiotics, which are classified as bacteriocidal if they kill bacteria, or bacteriostatic if they just prevent bacterial growth. There are many types of antibiotics and each class inhibits a process that is different in the pathogen from that found in the host. An example of how antibiotics produce selective toxicity are chloramphenicol and puromycin, which inhibit the bacterial ribosome, but not the structurally-different eukaryotic ribosome.[91] Infections can be prevented by antiseptic measures such as sterilizating the skin prior to piercing it with the needle of a syringe, and by proper care of indwelling catheters. Surgical and dental instruments are also sterilized to prevent contamination and infection by bacteria. Sanitizers and disinfectants are used to kill bacteria or other pathogens on surfaces to prevent contamination and further reduce the risk of infection.
Uses in technology and industry
Bacteria, often Lactobacillus in combination with yeasts and molds, have been used for thousands of years in the preparation of fermented foods such as cheese, pickles, soy sauce, sauerkraut, vinegar, wine, and yogurt.
The ability of bacteria to degrade a variety of organic compounds is remarkable and has been used in waste processing, and bioremediation. Bacteria capable of digesting the hydrocarbons in petroleum are often used to clean up oil spills.[92] Fertilizer was added to some of the beaches in Prince William Sound in an attempt to promote the growth of these naturally occurring bacteria after the infamous 1989 Exxon Valdez oil spill. These efforts were effective on beaches that were not too thickly covered in oil. Bacteria are also used for the bioremediation of industrial toxic wastes.[93] In the chemical industry, bacteria are most important in the production of enantiomerically pure chemicals for use as pharmaceuticals or agrochemicals.[94]
Bacteria can also be used in the place of pesticides in the biological pest control. This commonly uses Bacillus thuringiensis (also called BT), a Gram-positive, soil dwelling bacterium. This bacteria is used as a Lepidopteran-specific insecticide under trade names such as Dipel and Thuricide. Because of their specificity, these pesticides are regarded as Environmentally friendly, with little or no effect on humans, wildlife, pollinators, and most other beneficial insects.
Because of their ability to quickly grow, and the relative ease with which they can be manipulated, bacteria are the workhorses for the fields of molecular biology, genetics and biochemistry. By making mutations in bacterial DNA and examining the resulting phenotypes, scientists hcan determine the function of many different genes, enzymes and metabolic pathways. Lessons learned from bacteria are then be applied to more complex organisms. This aim of understanding the biochemistry of a cell reaches its most complex expression in the synthesis of huge amounts of enzyme kinetic and gene expression data into mathematical models of entire organisms. This is achievable in some well-studied bacteria, with models of Escherichia coli metabolism now being produced and tested.[95][96] This understanding of bacterial metabolism and genetics allows the use of biotechnology to bioengineer bacteria for the production of therapeutic proteins, such as insulin, growth factors or antibodies.[97][98]
See also
References
- ^ Woese C, Kandler O, Wheelis M (1990). "Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya". Proc Natl Acad Sci U S A. 87 (12): 4576–9. PMID 2112744.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Fredrickson J, Zachara J, Balkwill D, Kennedy D, Li S, Kostandarithes H, Daly M, Romine M, Brockman F (2004). "Geomicrobiology of high-level nuclear waste-contaminated vadose sediments at the hanford site, washington state". Appl Environ Microbiol. 70 (7): 4230–41. PMID 15240306.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Whitman W, Coleman D, Wiebe W (1998). "Prokaryotes: the unseen majority". Proc Natl Acad Sci U S A. 95 (12): 6578–83. PMID 9618454.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ a b Rappé M, Giovannoni S. "The uncultured microbial majority". Annu Rev Microbiol. 57: 369–94. PMID 14527284.
- ^ Sears C (2005). "A dynamic partnership: Celebrating our gut flora". Anaerobe. 11 (5): 247–51. PMID 16701579.
- ^ Ishige T, Honda K, Shimizu S (2005). "Whole organism biocatalysis". Curr Opin Chem Biol. 9 (2): 174–80. PMID 15811802.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Leeuwenhoek A (1753). "Part of a Letter from Mr Antony van Leeuwenhoek, concerning the Worms in Sheeps Livers, Gnats, and Animalcula in the Excrements of Frogs". Philosophical Transactions (1683-1775). 22: 509–518.
- ^ Leeuwenhoek A (1753). "Part of a Letter from Mr Antony van Leeuwenhoek, F. R. S. concerning Green Weeds Growing in Water, and Some Animalcula Found about Them". Philosophical Transactions (1683-1775). 23: 1304–1311.
- ^ Etymology of the word "bacteria" Online Etymology dictionary. Accessed 23 Nov 06
- ^ Pasteur's Papers on the Germ Theory LSU Law Center's Medical and Public Health Law Site, Historic Public Health Articles. Accessed 23 Nov 06
- ^ The Nobel Prize in Physiology or Medicine 1905 Nobelprize.org Accessed 22 Nov 06
- ^ O'Brien S, Goedert J (1996). "HIV causes AIDS: Koch's postulates fulfilled". Curr Opin Immunol. 8 (5): 613–8. PMID 8902385.
- ^ Schwartz R (2004). "Paul Ehrlich's magic bullets". N Engl J Med. 350 (11): 1079–80. PMID 15014180.
- ^ Biography of Paul Ehrlich Nobelprize.org Accessed 26 Nov 06
- ^ Woese C, Fox G (1977). "Phylogenetic structure of the prokaryotic domain: the primary kingdoms". Proc Natl Acad Sci U S A. 74 (11): 5088–90. PMID 270744.
- ^ Woese C, Kandler O, Wheelis M (1990). "Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya". Proc Natl Acad Sci U S A. 87 (12): 4576–9. PMID 2112744.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Schopf J (1994). "Disparate rates, differing fates: tempo and mode of evolution changed from the Precambrian to the Phanerozoic". Proc Natl Acad Sci U S A. 91 (15): 6735–42. PMID 8041691.
- ^ DeLong E, Pace N (2001). "Environmental diversity of bacteria and archaea". Syst Biol. 50 (4): 470–8. PMID 12116647.
- ^ Di Giulio M (2003). "The universal ancestor and the ancestor of bacteria were hyperthermophiles". J Mol Evol. 57 (6): 721–30. PMID 14745541.
- ^ Dyall S, Brown M, Johnson P (2004). "Ancient invasions: from endosymbionts to organelles". Science. 304 (5668): 253–7. PMID 15073369.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Rotte C, Henze K, Müller M, Martin W (2000). "Origins of hydrogenosomes and mitochondria". Curr Opin Microbiol. 3 (5): 481–6. PMID 11050446.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Schulz H, Jorgensen B. "Big bacteria". Annu Rev Microbiol. 55: 105–37. PMID 11544351.
- ^ Robertson J, Gomersall M, Gill P. (1975). "Mycoplasma hominis: growth, reproduction, and isolation of small viable cells". J Bacteriol. 124 (2): 1007–18. PMID 1102522.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Cabeen M, Jacobs-Wagner C (2005). "Bacterial cell shape". Nat Rev Microbiol. 3 (8): 601–10. PMID 16012516.
- ^ Young K (2006). "The selective value of bacterial shape". Microbiol Mol Biol Rev. 70 (3): 660–703. PMID 16959965.
- ^ Douwes K, Schmalzbauer E, Linde H, Reisberger E, Fleischer K, Lehn N, Landthaler M, Vogt T (2003). "Branched filaments no fungus, ovoid bodies no bacteria: Two unusual cases of mycetoma". J Am Acad Dermatol. 49 (2 Suppl Case Reports): S170-3. PMID 12894113.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Donlan R (2002). "Biofilms: microbial life on surfaces". Emerg Infect Dis. 8 (9): 881–90. PMID 12194761.
- ^ Branda S, Vik S, Friedman L, Kolter R (2005). "Biofilms: the matrix revisited". Trends Microbiol. 13 (1): 20–6. PMID 15639628.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ a b Davey M, O'toole G (2000). "Microbial biofilms: from ecology to molecular genetics". Microbiol Mol Biol Rev. 64 (4): 847–67. PMID 11104821.
- ^ Donlan RM, Costerton JW (2002). "Biofilms: survival mechanisms of clinically relevant microorganisms". Clin Microbiol Rev. 15 (2): 167–93. PMID 11932229.
- ^ Kaiser D. "Signaling in myxobacteria". Annu Rev Microbiol. 58: 75–98. PMID 15487930.
- ^ Harold F (1972). "Conservation and transformation of energy by bacterial membranes". Bacteriol Rev. 36 (2): 172–230. PMID 4261111.
- ^ Thanbichler M, Wang S, Shapiro L (2005). "The bacterial nucleoid: a highly organized and dynamic structure". J Cell Biochem. 96 (3): 506–21. PMID 15988757.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Poehlsgaard J, Douthwaite S (2005). "The bacterial ribosome as a target for antibiotics". Nat Rev Microbiol. 3 (11): 870–81. PMID 16261170.
- ^ Fuerst J (2005). "Intracellular compartmentation in planctomycetes". Annu Rev Microbiol. 59: 299–328. PMID 15910279.
- ^ Yeo M, Chater K (2005). "The interplay of glycogen metabolism and differentiation provides an insight into the developmental biology of Streptomyces coelicolor". Microbiology. 151 (Pt 3): 855–61. PMID 15758231.
- ^ Shiba T, Tsutsumi K, Ishige K, Noguchi T (2000). "Inorganic polyphosphate and polyphosphate kinase: their novel biological functions and applications". Biochemistry (Mosc). 65 (3): 315–23. PMID 10739474.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Walsby A (1994). "Gas vesicles". Microbiol Rev. 58 (1): 94–144. PMID 8177173.
- ^ van Heijenoort J (2001). "Formation of the glycan chains in the synthesis of bacterial peptidoglycan". Glycobiology. 11 (3): 25R–36R. PMID 11320055.
- ^ a b Koch A (2003). "Bacterial wall as target for attack: past, present, and future research". Clin Microbiol Rev. 16 (4): 673–87. PMID 14557293.
- ^ Hugenholtz P (2002). "Exploring prokaryotic diversity in the genomic era". Genome Biol. 3 (2): REVIEWS0003. PMID 11864374.
- ^ Engelhardt H, Peters J (1998). "Structural research on surface layers: a focus on stability, surface layer homology domains, and surface layer-cell wall interactions". J Struct Biol. 124 (2–3): 276–302. PMID 10049812.
- ^ Beveridge T, Pouwels P, Sára M, Kotiranta A, Lounatmaa K, Kari K, Kerosuo E, Haapasalo M, Egelseer E, Schocher I, Sleytr U, Morelli L, Callegari M, Nomellini J, Bingle W, Smit J, Leibovitz E, Lemaire M, Miras I, Salamitou S, Béguin P, Ohayon H, Gounon P, Matuschek M, Koval S (1997). "Functions of S-layers". FEMS Microbiol Rev. 20 (1–2): 99–149. PMID 9276929.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Kojima S, Blair D. "The bacterial flagellar motor: structure and function of a complex molecular machine". Int Rev Cytol. 233: 93–134. PMID 15037363.
- ^ Beachey E (1981). "Bacterial adherence: adhesin-receptor interactions mediating the attachment of bacteria to mucosal surface". J Infect Dis. 143 (3): 325–45. PMID 7014727.
- ^ Silverman P (1997). "Towards a structural biology of bacterial conjugation". Mol Microbiol. 23 (3): 423–9. PMID 9044277.
- ^ Stokes R, Norris-Jones R, Brooks D, Beveridge T, Doxsee D, Thorson L (2004). "The glycan-rich outer layer of the cell wall of Mycobacterium tuberculosis acts as an antiphagocytic capsule limiting the association of the bacterium with macrophages". Infect Immun. 72 (10): 5676–86. PMID 15385466.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Daffé M, Etienne G (1999). "The capsule of Mycobacterium tuberculosis and its implications for pathogenicity". Tuber Lung Dis. 79 (3): 153–69. PMID 10656114.
- ^ Nicholson W, Munakata N, Horneck G, Melosh H, Setlow P (2000). "Resistance of Bacillus endospores to extreme terrestrial and extraterrestrial environments". Microbiol Mol Biol Rev. 64 (3): 548–72. PMID 10974126.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Nicholson W, Fajardo-Cavazos P, Rebeil R, Slieman T, Riesenman P, Law J, Xue Y (2002). "Bacterial endospores and their significance in stress resistance". Antonie Van Leeuwenhoek. 81 (1–4): 27–32. PMID 12448702.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Vreeland R, Rosenzweig W, Powers D (2000). "Isolation of a 250 million-year-old halotolerant bacterium from a primary salt crystal". Nature. 407 (6806): 897–900. PMID 11057666.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Cano R, Borucki M (1995). "Revival and identification of bacterial spores in 25- to 40-million-year-old Dominican amber". Science. 268 (5213): 1060–4. PMID 7538699.
- ^ Nicholson W, Schuerger A, Setlow P (2005). "The solar UV environment and bacterial spore UV resistance: considerations for Earth-to-Mars transport by natural processes and human spaceflight". Mutat Res. 571 (1–2): 249–64. PMID 15748651.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Hatheway C (1990). "Toxigenic clostridia". Clin Microbiol Rev. 3 (1): 66–98. PMID 2404569.
- ^ Hellingwerf K, Crielaard W, Hoff W, Matthijs H, Mur L, van Rotterdam B (1994). "Photobiology of bacteria". Antonie Van Leeuwenhoek. 65 (4): 331–47. PMID 7832590.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Zumft W (1997). "Cell biology and molecular basis of denitrification". Microbiol Mol Biol Rev. 61 (4): 533–616. PMID 9409151.
- ^ Drake H, Daniel S, Küsel K, Matthies C, Kuhner C, Braus-Stromeyer S (1997). "Acetogenic bacteria: what are the in situ consequences of their diverse metabolic versatilities?". Biofactors. 6 (1): 13–24. PMID 9233536.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Dalton H (2005). "The Leeuwenhoek Lecture 2000 the natural and unnatural history of methane-oxidizing bacteria" (PDF). Philos Trans R Soc Lond B Biol Sci. 360 (1458): 1207–22. PMID 16147517.
- ^ Zehr J, Jenkins B, Short S, Steward G (2003). "Nitrogenase gene diversity and microbial community structure: a cross-system comparison". Environ Microbiol. 5 (7): 539–54. PMID 12823187.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Koch A (2002). "Control of the bacterial cell cycle by cytoplasmic growth". Crit Rev Microbiol. 28 (1): 61–77. PMID 12003041.
- ^ Eagon R. "Pseudomonas natriegens, a marine bacterium with a generation time of less than 10 minutes". J Bacteriol. 83: 736–7. PMID 13888946.
- ^ Paerl H, Fulton R, Moisander P, Dyble J. "Harmful freshwater algal blooms, with an emphasis on cyanobacteria". ScientificWorldJournal. 1: 76–113. PMID 12805693.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Challis G, Hopwood D. "Synergy and contingency as driving forces for the evolution of multiple secondary metabolite production by Streptomyces species". Proc Natl Acad Sci U S A. 100 Suppl 2: 14555–61. PMID 12970466.
- ^ Kooijman S, Auger P, Poggiale J, Kooi B (2003). "Quantitative steps in symbiogenesis and the evolution of homeostasis". Biol Rev Camb Philos Soc. 78 (3): 435–63. PMID 14558592.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Prats C, López D, Giró A, Ferrer J, Valls J (2006). "Individual-based modelling of bacterial cultures to study the microscopic causes of the lag phase". J Theor Biol. 241 (4): 939–53. PMID 16524598.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Hecker M, Völker U. "General stress response of Bacillus subtilis and other bacteria". Adv Microb Physiol. 44: 35–91. PMID 11407115.
- ^ Denamur E, Matic I (2006). "Evolution of mutation rates in bacteria". Mol Microbiol. 60 (4): 820–7. PMID 16677295.
- ^ Wright B (2004). "Stress-directed adaptive mutations and evolution". Mol Microbiol. 52 (3): 643–50. PMID 15101972.
- ^ Davison J (1999). "Genetic exchange between bacteria in the environment". Plasmid. 42 (2): 73–91. PMID 10489325.
- ^ Hastings P, Rosenberg S, Slack A (2004). "Antibiotic-induced lateral transfer of antibiotic resistance". Trends Microbiol. 12 (9): 401–4. PMID 15337159.
{{cite journal}}: CS1 maint: multiple names: authors list (link), - ^ a b c Bardy S, Ng S, Jarrell K (2003). "Prokaryotic motility structures". Microbiology. 149 (Pt 2): 295–304. PMID 12624192.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Wu M, Roberts J, Kim S, Koch D, DeLisa M (2006). "Collective bacterial dynamics revealed using a three-dimensional population-scale defocused particle tracking technique". Appl Environ Microbiol. 72 (7): 4987–94. PMID 16820497.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Lux R, Shi W (2004). "Chemotaxis-guided movements in bacteria". Crit Rev Oral Biol Med. 15 (4): 207–20. PMID 15284186.
- ^ Frankel R, Bazylinski D, Johnson M, Taylor B (1997). "Magneto-aerotaxis in marine coccoid bacteria". Biophys J. 73 (2): 994–1000. PMID 9251816.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Kaiser D. "Signaling in myxobacteria". Annu Rev Microbiol. 58: 75–98. PMID 15487930.
- ^ Sapp J (2005). "The prokaryote-eukaryote dichotomy: meanings and mythology". Microbiol Mol Biol Rev. 69 (2): 292–305. PMID 15944457.
- ^ Gram, C (1884). "Uber die isolirte Farbung der Schizomyceten in Schnitt- und Trockenpreparaten". Fortschr. Med. 2: 185–189.
- ^ Woods G, Walker D (1996). "Detection of infection or infectious agents by use of cytologic and histologic stains". Clin Microbiol Rev. 9 (3): 382–404. PMID 8809467.
- ^ Weinstein M (1994). "Clinical importance of blood cultures". Clin Lab Med. 14 (1): 9–16. PMID 8181237.
- ^ Thomson R, Bertram H (2001). "Laboratory diagnosis of central nervous system infections". Infect Dis Clin North Am. 15 (4): 1047–71. PMID 11780267.
- ^ Woese C, Kandler O, Wheelis M (1990). "Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya". Proc Natl Acad Sci U S A. 87 (12): 4576–9. PMID 2112744.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Stams A, de Bok F, Plugge C, van Eekert M, Dolfing J, Schraa G (2006). "Exocellular electron transfer in anaerobic microbial communities". Environ Microbiol. 8 (3): 371–82. PMID 16478444.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Barea J, Pozo M, Azcón R, Azcón-Aguilar C (2005). "Microbial co-operation in the rhizosphere". J Exp Bot. 56 (417): 1761–78. PMID 15911555.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ O'Hara A, Shanahan F (2006). "The gut flora as a forgotten organ". EMBO Rep. 7 (7): 688–93. PMID 16819463.
- ^ Zoetendal E, Vaughan E, de Vos W (2006). "A microbial world within us". Mol Microbiol. 59 (6): 1639–50. PMID 16553872.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Gorbach S (1990). "Lactic acid bacteria and human health". Ann Med. 22 (1): 37–41. PMID 2109988.
- ^ Salminen S, Gueimonde M, Isolauri E (2005). "Probiotics that modify disease risk". J Nutr. 135 (5): 1294–8. PMID 15867327.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Belland R, Ouellette S, Gieffers J, Byrne G (2004). "Chlamydia pneumoniae and atherosclerosis". Cell Microbiol. 6 (2): 117–27. PMID 14706098.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Fish D. "Optimal antimicrobial therapy for sepsis". Am J Health Syst Pharm. 59 Suppl 1: S13-9. PMID 11885408.
- ^ Heise E. "Diseases associated with immunosuppression". Environ Health Perspect. 43: 9–19. PMID 7037390.
- ^ Yonath A, Bashan A (2004). "Ribosomal crystallography: initiation, peptide bond formation, and amino acid polymerization are hampered by antibiotics". Annu Rev Microbiol. 58: 233–51. PMID 15487937.
- ^ Cohen Y (2002). "Bioremediation of oil by marine microbial mats". Int Microbiol. 5 (4): 189–93. PMID 12497184.
- ^ Neves L, Miyamura T, Moraes D, Penna T, Converti A. "Biofiltration methods for the removal of phenolic residues". Appl Biochem Biotechnol. 129–132: 130–52. PMID 16915636.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ Liese A, Filho M (1999). "Production of fine chemicals using biocatalysis". Curr Opin Biotechnol. 10 (6): 595–603. PMID 10600695.
- ^ Almaas E, Kovacs B, Vicsek T, Oltvai ZN, Barabasi AL. Global organization of metabolic fluxes in the bacterium Escherichia coli. Nature. 2004 Feb 26;427(6977):839-43. PMID 14985762
- ^ Reed JL, Vo TD, Schilling CH, Palsson BO. An expanded genome-scale model of Escherichia coli K-12 (iJR904 GSM/GPR). Genome Biol. 2003;4(9):R54. PMID 12952533
- ^ Walsh G (2005). "Therapeutic insulins and their large-scale manufacture". Appl Microbiol Biotechnol. 67 (2): 151–9. PMID 15580495.
- ^ Graumann K, Premstaller A (2006). "Manufacturing of recombinant therapeutic proteins in microbial systems". Biotechnol J. 1 (2): 164–86. PMID 16892246.
Further reading
- Alcamo, I. Edward. Fundamentals of Microbiology. 5th ed. Menlo Park, California: Benjamin Cumming, 1997.
- Atlas, Ronald M. Principles of Microbiology. St. Louis, Missouri: Mosby, 1995.
- Holt, John.G. Bergey's Manual of Determinative Bacteriology. 9th ed. Baltimore, Maryland: Williams and Wilkins, 1994.
- Hugenholtz P, Goebel BM, Pace NR (1998). "Impact of culture-independent studies on the emerging phylogenetic view of bacterial diversity". J Bacteriol. 180 (18): 4765–74. PMID 9733676.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - Koshland, Daniel E., Jr. (1977). "A Response Regulator Model in a Simple Sensory System". Science. 196: 1055–1063. PMID 870969.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - Stanier, R.Y., J. L. Ingraham, M. L. Wheelis, and P. R. Painter. General Microbiology. 5th ed. Upper Saddle River, New Jersey: Prentice Hall, 1986.
External links
- Bacterial Nomenclature Up-To-Date from DSMZ
- Bacterial Growth and Cell Wall (Ger)
- Microminds
- The largest bacteria
- Tree of Life
- Videos of bacteria swimming and tumbling, use of optical tweezers and other fine videos.
- Planet of the Bacteria by Stephen Jay Gould
- Major Groups of Prokaryotes
- Bitter Resistance by Bruce Sterling
- On-line text book on bacteriology
- cyanobacteria in lichens
