In biology, scaffold proteins are crucial regulators of many key signalling pathways. Although scaffolds are not strictly defined in function, they are known to interact and/or bind with multiple members of a signalling pathway, tethering them into complexes. In such pathways, they regulate signal transduction and help localize pathway components (organized in complexes) to specific areas of the cell such as the plasma membrane, the cytoplasm, the nucleus, the Golgi, endosomes, and the mitochondria.
History[edit]
The first signaling scaffold protein discovered was the Ste5 protein from the yeast Saccharomyces cerevisiae. Three distinct domains of Ste5 were shown to associate with the protein kinases Ste11, Ste7, and Fus3 to form a multikinase complex.[2]
Function[edit]
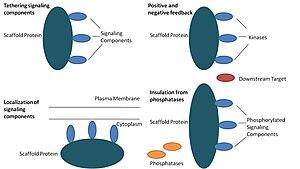
Scaffold proteins act in at least four ways: tethering signaling components, localizing these components to specific areas of the cell, regulating signal transduction by coordinating positive and negative feedback signals, and insulating correct signaling proteins from competing proteins.[1]
Tethering signaling components[edit]
This particular function is considered a scaffold's most basic function. Scaffolds assemble signaling components of a cascade into complexes. This assembly may be able to enhance signaling specificity by preventing unnecessary interactions between signaling proteins, and enhance signaling efficiency by increasing the proximity and effective concentration of components in the scaffold complex. A common example of how scaffolds enhance specificity is a scaffold that binds a protein kinase and its substrate, thereby ensuring specific kinase phosphorylation. Additionally, some signaling proteins require multiple interactions for activation and scaffold tethering may be able to convert these interactions into one interaction that results in multiple modifications.[3][4] Scaffolds may also be catalytic as interaction with signaling proteins may result in allosteric changes of these signaling components.[5] Such changes may be able to enhance or inhibit the activation of these signaling proteins. An example is the Ste5 scaffold in the mitogen-activated protein kinase (MAPK) pathway. Ste5 has been proposed to direct mating signaling through the Fus3 MAPK by catalytically unlocking this particular kinase for activation by its MAPKK Ste7.[6]
Localization of signaling components in the cell[edit]
Scaffolds localize the signaling reaction to a specific area in the cell, a process that could be important for the local production of signaling intermediates. A particular example of this process is the scaffold, A-kinase anchor proteins (AKAPs), which target cyclic AMP-dependent protein kinase (PKA) to various sites in the cell.[7] This localization is able to locally regulate PKA and results in the local phosphorylation by PKA of its substrates.
Coordinating positive and negative feedback[edit]
Many hypotheses about how scaffolds coordinate positive and negative feedback come from engineered scaffolds and mathematical modeling. In three-kinase signaling cascades, scaffolds bind all three kinases, enhancing kinase specificity and restricting signal amplification by limiting kinase phosphorylation to only one downstream target.[3][8][9] These abilities may be related to stability of the interaction between the scaffold and the kinases, the basal phosphatase activity in the cell, scaffold location, and expression levels of the signaling components.[3][8]
Insulating correct signaling proteins from inactivation[edit]
Signaling pathways are often inactivated by enzymes that reverse the activation state and/or induce the degradation of signaling components. Scaffolds have been proposed to protect activated signaling molecules from inactivation and/or degradation. Mathematical modeling has shown that kinases in a cascade without scaffolds have a higher probability of being dephosphorylated by phosphatases before they are even able to phosphorylate downstream targets.[8] Furthermore, scaffolds have been shown to insulate kinases from substrate- and ATP-competitive inhibitors.[10]
Scaffold protein summary[edit]
| Scaffold Proteins | Pathway | Potential Functions | Description |
|---|---|---|---|
| KSR | MAPK | Assembly and localization of the RAS-ERK pathway | One of the best studied signaling pathways in biology is the RAS-ERK pathway in which the RAS G-protein activates the MAPKKK RAF, which activates the MAPKK MEK1 (MAPK/ERK kinase 1), which then activates the MAPK ERK. Several scaffold proteins have been identified to be involved in this pathway and other similar MAPK pathways. One such scaffold protein is KSR, which is the most probable equivalent of the well-studied yeast MAPK scaffold protein Ste5.[11] It is a positive regulator of the pathway and binds many proteins in the pathway, including all three kinases in the cascade.[6] KSR has been shown to be localized to the plasma membrane during cell activation, thereby playing a role in assembling the components of the ERK pathway and in localizing activated ERK to the plasma membrane.[12] |
| MEKK1 | MAPK | Assembly and localization of the death receptor signalosome | Other scaffold proteins include B-cell lymphoma 10 (BCL-10) and MEK kinase 1 (MEKK1), which have roles in the JUN N-terminal kinase (JNK) pathway. |
| BCL-10 | MAPK | Assembly and specificity of JNK | |
| AKAP | PKA Pathways | Coordination of phosphorylation by PKA onto downstream targets | This family of proteins is only structurally related in their ability to bind the regulatory subunit of PKA but can otherwise bind a very diverse set of enzymes and substrates |
| AHNAK-1 | Calcium signaling | Assembly and localization of calcium channels | Calcium signaling is essential for the proper function of immune cells. Recent studies have shown that the scaffold protein, AHNAK1, is important for efficient calcium signaling and NFAT activation in T cells through its ability to properly localize calcium channels at the plasma membrane [14]. In non-immune cells, AHNAK1 has also been shown to bind calcium channels with phospholipase Cγ (PLC-γ) and PKC.[1] Calcium binding proteins often quench much of the entering calcium, so linking these calcium effectors may be especially important when signals are induced by a weak calcium influx. |
| HOMER | Calcium signaling | Inhibition of NFAT activation | Another example of a scaffold protein that modulates calcium signaling is proteins of the HOMER family. The HOMER proteins have been shown to compete with calcineurin to bind to the N terminus of NFAT in activated T cells.[13] Through this competition, the HOMER proteins are able to reduce NFAT activation, which also reduces the production of the IL-2 cytokine.[13] In contrast, HOMER proteins have also been shown to positively regulate calcium signaling in neurons by linking the glutamate receptor with triphosphate receptors in the endoplasmic reticulum.[14] |
| Pellino | Innate Immune Signaling | Assembly of the TLR signalosome | Evidence exists that Pellino proteins function as scaffold proteins in the important innate immune signaling pathway, the Toll-like receptor (TLR) pathway. Much Pellino function is speculation; however, Pellino proteins can associate with IRAK1, TRAF6, and TAK1 following IL-1R activation, indicating that they may assemble and localize components of the TLR pathway near its receptor.[15][16] |
| NLRP | Innate Immune Signaling | Assembly of the inflammasome | The NLR family is a highly conserved and large family of receptors involved in innate immunity. The NLRP (NLR family, pyrine domain-containing) family of receptors function as scaffolds by assembling the inflammasome, a complex that leads to the secretion of pro-inflammatory cytokines such as IL-18 and IL-1β.[17] |
| DLG1 | T-cell receptor signaling | Assembly and localization of TCR signaling molecules, activation of p38 | DLG1 is highly conserved in immune cells and is important for T-cell activation in the periphery. It is recruited to the immunological synapse and links the ζ-chain of the T-cell receptor (TCR) to CBL, WASP, p38, LCK, VAV1, and ZAP70.[18][19][20][21] This data suggests that DLG1 plays a role in linking TCR signaling machinery with cytoskeleton regulators and also suggests a role in alternatively activating the p38 pathway. However, it is unclear to whether DLG1 positively or negatively regulates T-cell activation. |
| Spinophilin | Dendritic cell signaling | Assembly of DC immunological-synapse proteins | Spinophilin is involved in dendritic cell function specifically in the formation of immunological synapses. Spinophilin is recruited to the synapse following dendritic cell contact with a T cell. This recruitment seems to be important because without spinophilin, dendritic cells cannot activate T cells in vitro or in vivo.[22] How spinophilin facilitates antigen presentation in this case is still unknown though it is possible that spinophilin regulates the duration of cell contact in the synapse or regulates the recycling of co-stimulatory molecules in the cell like MHC molecules.[1] |
| Plant FLU regulatory protein[23] | Coordination of negative feedback during protochlorophyllide biosynthesis. | Assembly and localization of the pathway that turns the synthesis of highly toxic protochlorophyllide, a precursor of chlorophyll. | Synthesis of protochlorophyllide must be strictly regulated as its conversion into chlorophyll requires light. FLU regulatory protein is located in thylakoid membrane and only contains several protein-protein interaction sites without catalytic activity. Mutants lacking this protein overaccumulate protochlorophyllide in the darkness. The interaction partners are unknown. The protein underwent simplification during evolution. |
Huntingtin protein[edit]
Huntingtin protein co-localizes with ATM repair protein at sites of DNA damage.[24] Huntingtin is a scaffolding protein in the ATM oxidative DNA damage response complex.[24] Huntington's disease patients with aberrant huntingtin protein are deficient in repair of oxidative DNA damage. Oxidative DNA damage appears to underlie Huntington's disease pathogenesis.[25] Huntington's disease is likely caused by the dysfunction of mutant huntingtin scaffold protein in DNA repair leading to increased oxidative DNA damage in metabolically active cells.[24]
DNA repair[edit]
SPIDR (scaffold protein involved in DNA repair) regulates the stability or assembly of RAD51 and DMC1 on single-stranded DNA.[26] RAD51 and DMC1 are recombinases that act during mammalian meiosis to mediate strand exchange during the repair of DNA double-strand breaks by homologous recombination.[26]
Other usage of the term Scaffold Protein[edit]
On some other instances in biology (not necessarily about cell signaling), the term "Scaffold protein" is used in a broader sense, where a protein holds several things together for any purpose.
- In chromosome folding
- Chromosome scaffold has important role to hold the chromatin into compact chromosome. Chromosome scaffold is made of proteins including condensin, topoisomerase IIα and kinesin family member 4 (KIF4)[27] Chromosome scaffold constituent proteins are also called scaffold protein.
- In enzymatic reaction
- Large multifunctional enzymes that performs a series or chain of reaction in a common pathway, sometimes called scaffold proteins.[28] such as Pyruvate dehydrogenase.
- In molecule shape formation
- An enzyme or structural protein that holds several molecules together to hold them in proper spatial arrangement, such as Iron sulphur cluster scaffold proteins.[29][30]
- Structural scaffold
- In cytoskeleton and ECM, the molecules provide mechanical scaffold. Such as type 4 collagen[31]
References[edit]
- ^ a b c d Shaw, Andrey S.; Filbert, Erin L. (January 2009). "Scaffold proteins and immune-cell signalling". Nature Reviews Immunology. 9 (1): 47–56. doi:10.1038/nri2473. PMID 19104498. S2CID 13443447.
- ^ Choi, Kang-Yell; Satterberg, Brett; Lyons, David M.; Elion, Elaine A. (August 1994). "Ste5 tethers multiple protein kinases in the MAP kinase cascade required for mating in S. cerevisiae". Cell. 78 (3): 499–512. doi:10.1016/0092-8674(94)90427-8. PMID 8062390. S2CID 20541545.
- ^ a b c Levchenko, Andre; Bruck, Jehoshua; Sternberg, Paul W. (23 May 2000). "Scaffold proteins may biphasically affect the levels of mitogen-activated protein kinase signaling and reduce its threshold properties". Proceedings of the National Academy of Sciences. 97 (11): 5818–5823. Bibcode:2000PNAS...97.5818L. doi:10.1073/pnas.97.11.5818. PMC 18517. PMID 10823939.
- ^ Ferrell, James E. (3 October 2000). "What Do Scaffold Proteins Really Do?". Science Signaling. 2000 (52): pe1. doi:10.1126/stke.522000pe1. S2CID 219192522.
- ^ Burack, W Richard; Shaw, Andrey S (April 2000). "Signal transduction: hanging on a scaffold". Current Opinion in Cell Biology. 12 (2): 211–216. doi:10.1016/S0955-0674(99)00078-2. PMID 10712921.
- ^ a b Good, Matthew; Tang, Grace; Singleton, Julie; Reményi, Attila; Lim, Wendell A. (March 2009). "The Ste5 Scaffold Directs Mating Signaling by Catalytically Unlocking the Fus3 MAP Kinase for Activation". Cell. 136 (6): 1085–1097. doi:10.1016/j.cell.2009.01.049. PMC 2777755. PMID 19303851.
- ^ Wong, Wei; Scott, John D. (December 2004). "AKAP signalling complexes: focal points in space and time". Nature Reviews Molecular Cell Biology. 5 (12): 959–970. doi:10.1038/nrm1527. PMID 15573134. S2CID 15268680.
- ^ a b c Locasale, Jason W.; Shaw, Andrey S.; Chakraborty, Arup K. (14 August 2007). "Scaffold proteins confer diverse regulatory properties to protein kinase cascades". Proceedings of the National Academy of Sciences. 104 (33): 13307–13312. Bibcode:2007PNAS..10413307L. doi:10.1073/pnas.0706311104. PMC 1948937. PMID 17686969. S2CID 8907943.
- ^ Uhlik, Mark T; Abell, Amy N; Cuevas, Bruce D; Nakamura, Kazuhiro; Johnson, Gary L (1 December 2004). "Wiring diagrams of MAPK regulation by MEKK1, 2, and 3". Biochemistry and Cell Biology. 82 (6): 658–663. doi:10.1139/o04-114. PMID 15674433.
- ^ Greenwald, Eric C.; Redden, John M.; Dodge-Kafka, Kimberly L.; Saucerman, Jeffrey J. (24 January 2014). "Scaffold State Switching Amplifies, Accelerates, and Insulates Protein Kinase C Signaling". Journal of Biological Chemistry. 289 (4): 2353–2360. doi:10.1074/jbc.M113.497941. PMC 3900978. PMID 24302730.
- ^ Clapéron, A.; Therrien, M. (May 2007). "KSR and CNK: two scaffolds regulating RAS-mediated RAF activation". Oncogene. 26 (22): 3143–3158. doi:10.1038/sj.onc.1210408. PMID 17496912. S2CID 31061333.
- ^ Müller, Jürgen; Ory, Stéphane; Copeland, Terry; Piwnica-Worms, Helen; Morrison, Deborah K. (November 2001). "C-TAK1 Regulates Ras Signaling by Phosphorylating the MAPK Scaffold, KSR1". Molecular Cell. 8 (5): 983–993. doi:10.1016/S1097-2765(01)00383-5. PMID 11741534.
- ^ a b Huang, Guo N.; Huso, David L.; Bouyain, Samuel; Tu, Jianchen; McCorkell, Kelly A.; May, Michael J.; Zhu, Yuwen; Lutz, Michael; Collins, Samuel; Dehoff, Marlin; Kang, Shin; Whartenby, Katharine; Powell, Jonathan; Leahy, Daniel; Worley, Paul F. (25 January 2008). "NFAT Binding and Regulation of T Cell Activation by the Cytoplasmic Scaffolding Homer Proteins". Science. 319 (5862): 476–481. Bibcode:2008Sci...319..476H. doi:10.1126/science.1151227. PMC 3602998. PMID 18218901.
- ^ Xiao, Bo; Cheng Tu, Jian; Worley, Paul F (June 2000). "Homer: a link between neural activity and glutamate receptor function". Current Opinion in Neurobiology. 10 (3): 370–374. doi:10.1016/S0959-4388(00)00087-8. PMID 10851183. S2CID 8699597.
- ^ Jiang, Zhengfan; Johnson, H. Jan; Nie, Huiqing; Qin, Jinzhong; Bird, Timothy A.; Li, Xiaoxia (28 March 2003). "Pellino 1 Is Required for Interleukin-1 (IL-1)-mediated Signaling through Its Interaction with the IL-1 Receptor-associated Kinase 4 (IRAK4)-IRAK-Tumor Necrosis Factor Receptor-associated Factor 6 (TRAF6) Complex". Journal of Biological Chemistry. 278 (13): 10952–10956. doi:10.1074/jbc.M212112200. PMID 12496252. S2CID 10165785.
- ^ Yu, Kang-Yeol; Kwon, Hyung-Joo; Norman, David A. M.; Vig, Eva; Goebl, Mark G.; Harrington, Maureen A. (15 October 2002). "Cutting Edge: Mouse Pellino-2 Modulates IL-1 and Lipopolysaccharide Signaling". The Journal of Immunology. 169 (8): 4075–4078. doi:10.4049/jimmunol.169.8.4075. PMID 12370331. S2CID 25317655.
- ^ Pétrilli, Virginie; Dostert, Catherine; Muruve, Daniel A; Tschopp, Jürg (December 2007). "The inflammasome: a danger sensing complex triggering innate immunity". Current Opinion in Immunology. 19 (6): 615–622. doi:10.1016/j.coi.2007.09.002. PMID 17977705.
- ^ Xavier, Ramnik; Rabizadeh, Shahrooz; Ishiguro, Kazuhiro; Andre, Niko; Ortiz, J. Bernabe; Wachtel, Heather; Morris, David G.; Lopez-Ilasaca, Marco; Shaw, Albert C.; Swat, Wojciech; Seed, Brian (19 July 2004). "Discs large (Dlg1) complexes in lymphocyte activation". Journal of Cell Biology. 166 (2): 173–178. doi:10.1083/jcb.200309044. PMC 2172307. PMID 15263016.
- ^ Hanada, Toshihiko; Lin, Lunhui; Chandy, K. George; Oh, S. Steven; Chishti, Athar H. (24 October 1997). "Human Homologue of the Drosophila Discs Large Tumor Suppressor Binds to p56 lck Tyrosine Kinase and Shaker Type Kv1.3 Potassium Channel in T Lymphocytes". Journal of Biological Chemistry. 272 (43): 26899–26904. doi:10.1074/jbc.272.43.26899. PMID 9341123. S2CID 23446334.
- ^ Round, June L.; Humphries, Lisa A.; Tomassian, Tamar; Mittelstadt, Paul; Zhang, Min; Miceli, M. Carrie (February 2007). "Scaffold protein Dlgh1 coordinates alternative p38 kinase activation, directing T cell receptor signals toward NFAT but not NF-κB transcription factors". Nature Immunology. 8 (2): 154–161. doi:10.1038/ni1422. PMID 17187070. S2CID 11906543.
- ^ Round, June L.; Tomassian, Tamar; Zhang, Min; Patel, Viresh; Schoenberger, Stephen P.; Miceli, M. Carrie (7 February 2005). "Dlgh1 coordinates actin polymerization, synaptic T cell receptor and lipid raft aggregation, and effector function in T cells". Journal of Experimental Medicine. 201 (3): 419–430. doi:10.1084/jem.20041428. PMC 2213022. PMID 15699074.
- ^ Bloom, Ona; Unternaehrer, Julia J.; Jiang, Aimin; Shin, Jeong-Sook; Delamarre, Lélia; Allen, Patrick; Mellman, Ira (21 April 2008). "Spinophilin participates in information transfer at immunological synapses". Journal of Cell Biology. 181 (2): 203–211. doi:10.1083/jcb.200711149. PMC 2315669. PMID 18411312. S2CID 1717736.
- ^ Meskauskiene, Rasa; Nater, Mena; Goslings, David; Kessler, Felix; Camp, Roel op den; Apel, Klaus (23 October 2001). "FLU: A negative regulator of chlorophyll biosynthesis in Arabidopsis thaliana". Proceedings of the National Academy of Sciences. 98 (22): 12826–12831. Bibcode:2001PNAS...9812826M. doi:10.1073/pnas.221252798. PMC 60138. PMID 11606728.
- ^ a b c Maiuri, Tamara; Mocle, Andrew J.; Hung, Claudia L.; Xia, Jianrun; van Roon-Mom, Willeke M. C.; Truant, Ray (25 December 2016). "Huntingtin is a scaffolding protein in the ATM oxidative DNA damage response complex". Human Molecular Genetics. 26 (2): 395–406. doi:10.1093/hmg/ddw395. PMID 28017939.
- ^ Ayala-Peña, Sylvette (September 2013). "Role of oxidative DNA damage in mitochondrial dysfunction and Huntington's disease pathogenesis". Free Radical Biology and Medicine. 62: 102–110. doi:10.1016/j.freeradbiomed.2013.04.017. PMC 3722255. PMID 23602907.
- ^ a b Huang T, Wu X, Wang S, Bao Z, Wan Y, Wang Z, Li M, Yu X, Lv Y, Liu Z, Chen X, Chan WY, Gao F, Lu G, Chen ZJ, Liu H. SPIDR is required for homologous recombination during mammalian meiosis. Nucleic Acids Res. 2023 May 8;51(8):3855-3868. doi: 10.1093/nar/gkad154. PMID: 36938872; PMCID: PMC10164582
- ^ Poonperm, Rawin; Takata, Hideaki; Hamano, Tohru; Matsuda, Atsushi; Uchiyama, Susumu; Hiraoka, Yasushi; Fukui, Kiichi (1 July 2015). "Chromosome Scaffold is a Double-Stranded Assembly of Scaffold Proteins". Scientific Reports. 5 (1): 11916. Bibcode:2015NatSR...511916P. doi:10.1038/srep11916. PMC 4487240. PMID 26132639.
- ^ molecular cell biology by Lodish[full citation needed]
- ^ Ayala-Castro, Carla; Saini, Avneesh; Outten, F. Wayne (2008). "Fe-S Cluster Assembly Pathways in Bacteria". Microbiology and Molecular Biology Reviews. 72 (1): 110–125. doi:10.1128/MMBR.00034-07. PMC 2268281. PMID 18322036.
- ^ Adrover, Miquel; Howes, Barry D.; Iannuzzi, Clara; Smulevich, Giulietta; Pastore, Annalisa (1 June 2015). "Anatomy of an iron-sulfur cluster scaffold protein: Understanding the determinants of [2Fe–2S] cluster stability on IscU". Biochimica et Biophysica Acta (BBA) - Molecular Cell Research. 1853 (6): 1448–1456. doi:10.1016/j.bbamcr.2014.10.023. PMID 25447544.
- ^ Molecular Cell Biology by Lodish et al. edition 5[page needed]