| OTC | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
 | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | OTC, OCTD, ornithine carbamoyltransferase, ornithine transcarbamylase, OTCD | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | OMIM: 300461 MGI: 97448 HomoloGene: 446 GeneCards: OTC | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
Ornithine transcarbamylase (OTC) (also called ornithine carbamoyltransferase) is an enzyme (EC 2.1.3.3) that catalyzes the reaction between carbamoyl phosphate (CP) and ornithine (Orn) to form citrulline (Cit) and phosphate (Pi). There are two classes of OTC: anabolic and catabolic. This article focuses on anabolic OTC. Anabolic OTC facilitates the sixth step in the biosynthesis of the amino acid arginine in prokaryotes.[5] In contrast, mammalian OTC plays an essential role in the urea cycle, the purpose of which is to capture toxic ammonia and transform it into urea, a less toxic nitrogen source, for excretion.
Reaction mechanism[edit]

Structure[edit]
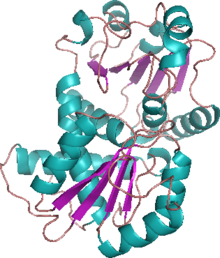
OTC is a trimeric protein. There are three active sites of the protein which are located at the cleft between the monomers. The carbamoyl phosphate binding domain resides on the N-terminal end of each monomer, while the C-terminal end contains the binding domain for ornithine. Both binding domains have a similar structural pattern with a central parallel β-pleated sheet bordered by α-helices and loops.[7] In addition to the binding domains, OTCs have SMG loops. These swing to close the binding site once both substrates have bound. SMG stands for the conserved amino acid motif of Ser-Met-Gly. Upon closure, these residues interact with L-ornithine. The binding of CP induces a global conformational change, while the binding of L-ornithine only induces movement of the SMG loop to close and isolate the activation site.[8]

Active site[edit]

Ser-Thr-Arg-Thr-Arg motif from one subunit and a His from the neighboring subunit both interact with the phosphate group of CP for binding. Binding the primary nitrogen of CP are residues Gln, Cys, and Arg. The carbonyl oxygen of CP is bound by residues Thr, Arg, and His.[10]
Amino acid composition[edit]
Plant OTCs have the largest difference from other OTCs. There are 50 to 70% less Leu residues, while there are twice as many Arg residues. The number of subunits in OTCs vary from 322 to 340 residues. Animals have the highest density of Leu. This residue breakdown causes a pI for the animal enzyme of 6.8 while the plant enzyme has a pI of 7.6.[11] Rat, bovine, and human OTC have the same C terminal residue of phenylalanine. Their N-terminal residues on the other hand differ. Rat ends with Ser, bovine with aspartate, and human with glycine.[12][13]
Genomics[edit]
The human OTC gene is located on the short arm of chromosome X (Xp21.1). The gene is located in the Watson (plus) strand and is 73 kbases in length. The open reading frame of 1,062 nucleotides is disbursed between 10 exons and nine introns. The encoded protein is 354 amino acids long with a predicted molecular weight of 39.935 kD. Postranscriptional modification leaves the mature peptide with 322 amino acids and a weight of 36.1 kD.[14] The protein is located in the mitochondrial matrix. In mammals, OTC is expressed in the liver and small intestinal mucosa.
Human mutations[edit]
341 mutations in human OTC have been reported. At least 259 of these mutations are considered to be disease-causing mutations.[15] 149 of these mutations are known to cause onset of hyperammonemia during the first weeks of life. 70 manifest as hyperammonemia in male patients later in life. Most of the mutations occur in known functional motifs, such as the SMG loop or CP binding domains.[16]
Deficiency[edit]
Mutations in the OTC gene can cause Ornithine Transcarbamylase deficiency. It is classified as a urea cycle disorder due to the fact that without proper OTC function ammonia starts to accumulate in the blood. Accumulation of ammonia in the blood is known as hyperammonemia. Although toxic in excess, ammonia is a nitrogen source for the body. Therefore increased ammonia will also increase levels of the nitrogen-containing non-essential amino acids glutamate, glutamine, and alanine. Levels of carbamoyl phosphate (CP) will begin to drop as urea nitrogen levels in the blood decrease. This will cause CP to be diverted to the uridine monophosphate synthetic pathway. Orotic acid is a product of this pathway. Increased levels of orotic acid in urine can be an indicator that a patient is suffering from a disorder linked to hyperammonemia.
OTC deficiency manifests in both early and late onset forms.
Early onset[edit]
Early onset is seen in newborns. The symptoms of a urea cycle disorder are often not seen until the child is at home and may not be recognized in a timely manner by the family and primary care physician. Symptoms in young children with hyperammonemia are non-specific: not willing to eat, problems with breathing, body temperature, seizures, unusual body movements (twitches) and somnolence.[17] As ammonia build up continues, symptoms progress from somnolence to lethargy potentially ending in a coma. Abnormal posturing (uncontrolled movement) and encephalopathy (brain damage) are often related to the degree of central nervous system swelling and pressure upon the brainstem. About 50% of neonates with severe hyperammonemia have seizures.
Late onset[edit]
In milder (or partial) urea cycle enzyme deficiencies, ammonia accumulation may be triggered by illness or stress at almost any time of life, resulting in multiple mild elevations of plasma ammonia concentration [Bourrier et al. 1988]. Patients with partial enzyme deficiencies may have a delay of symptoms for months or years. Indicators that you maybe suffering from OTC deficiency or a urea cycle disorder include "episodes of delirium, erratic behavior, or reduced consciousness, headaches, vomiting, aversion to foods high in protein, and seizures."[18]
Treatment[edit]
A potential treatment for the high ammonia levels is to give sodium benzoate, which combines with glycine to produce hippurate, at the same time removing an ammonium group. Biotin also plays an important role in the functioning of the OTC enzyme[19] and has been shown to reduce ammonia intoxication in animal experiments. Additionally, the use of whole-body therapeutic hypothermia (TH) has been proposed and studied as a treatment. TH is thought to increase the effectiveness of dialysis to extract ammonia from the body.[20][21]
References[edit]
- ^ a b c GRCh38: Ensembl release 89: ENSG00000036473 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000031173 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ Cunin R, Glansdorff N, Piérard A, Stalon V (September 1986). "Biosynthesis and metabolism of arginine in bacteria". Microbiological Reviews. 50 (3): 314–52. doi:10.1128/mr.50.3.314-352.1986. PMC 373073. PMID 3534538.
- ^ Langley DB, Templeton MD, Fields BA, Mitchell RE, Collyer CA (June 2000). "Mechanism of inactivation of ornithine transcarbamoylase by Ndelta -(N'-Sulfodiaminophosphinyl)-L-ornithine, a true transition state analogue? Crystal structure and implications for catalytic mechanism". The Journal of Biological Chemistry. 275 (26): 20012–9. doi:10.1074/jbc.M000585200. PMID 10747936.
- ^ Sankaranarayanan R, Cherney MM, Cherney LT, Garen CR, Moradian F, James MN (January 2008). "The crystal structures of ornithine carbamoyltransferase from Mycobacterium tuberculosis and its ternary complex with carbamoyl phosphate and L-norvaline reveal the enzyme's catalytic mechanism". Journal of Molecular Biology. 375 (4): 1052–63. doi:10.1016/j.jmb.2007.11.025. PMID 18062991.
- ^ Ha Y, McCann MT, Tuchman M, Allewell NM (September 1997). "Substrate-induced conformational change in a trimeric ornithine transcarbamoylase". Proceedings of the National Academy of Sciences of the United States of America. 94 (18): 9550–5. Bibcode:1997PNAS...94.9550H. doi:10.1073/pnas.94.18.9550. PMC 23215. PMID 9275160.
- ^ a b PDB: 1C9Y; Shi D, Morizono H, Aoyagi M, Tuchman M, Allewell NM (June 2000). "Crystal structure of human ornithine transcarbamylase complexed with carbamoyl phosphate and L-norvaline at 1.9 A resolution". Proteins. 39 (4): 271–7. doi:10.1002/(SICI)1097-0134(20000601)39:4<271::AID-PROT10>3.0.CO;2-E. PMID 10813810. S2CID 22697980.
- ^ Shi D, Morizono H, Yu X, Tong L, Allewell NM, Tuchman M (March 2001). "Human ornithine transcarbamylase: crystallographic insights into substrate recognition and conformational changes". The Biochemical Journal. 354 (Pt 3): 501–9. doi:10.1042/bj3540501. PMC 1221681. PMID 11237854.
- ^ Slocum RD, Richardson DP (1991-05-01). "Purification and characterization of ornithine transcarbamylase from pea (Pisum sativum L.)". Plant Physiology. 96 (1): 262–8. doi:10.1104/pp.96.1.262. PMC 1080743. PMID 11538003.
- ^ Kalousek F, François B, Rosenberg LE (June 1978). "Isolation and characterization of ornithine transcarbamylase from normal human liver". The Journal of Biological Chemistry. 253 (11): 3939–44. doi:10.1016/S0021-9258(17)34781-6. PMID 25896.
- ^ Lusty CJ, Jilka RL, Nietsch EH (October 1979). "Ornithine transcarbamylase of rat liver. Kinetic, physical, and chemical properties". The Journal of Biological Chemistry. 254 (20): 10030–6. doi:10.1016/S0021-9258(19)86668-1. PMID 489581.
- ^ Horwich AL, Kalousek F, Fenton WA, Pollock RA, Rosenberg LE (February 1986). "Targeting of pre-ornithine transcarbamylase to mitochondria: definition of critical regions and residues in the leader peptide". Cell. 44 (3): 451–9. doi:10.1016/0092-8674(86)90466-6. PMID 3943133. S2CID 23799662.
- ^ Šimčíková D, Heneberg P (December 2019). "Refinement of evolutionary medicine predictions based on clinical evidence for the manifestations of Mendelian diseases". Scientific Reports. 9 (1): 18577. Bibcode:2019NatSR...918577S. doi:10.1038/s41598-019-54976-4. PMC 6901466. PMID 31819097.
- ^ Yamaguchi S, Brailey LL, Morizono H, Bale AE, Tuchman M (July 2006). "Mutations and polymorphisms in the human ornithine transcarbamylase (OTC) gene". Human Mutation. 27 (7): 626–32. doi:10.1002/humu.20339. PMID 16786505. S2CID 26009099.
- ^ "Ornithine transcarbamylase deficiency". Genetics Home Reference. National Library of Medicine, U.S. Department of Health & Human Services. Retrieved 2019-03-03.
- ^ "Ornithine transcarbamylase deficiency". Genetic and Rare Diseases Information Center (GARD) – an NCATS Program. National Institutes of Health, U.S. Department of Health & Human Services. Retrieved 2019-03-03.
- ^ Nagamine T, Saito S, Kaneko M, Sekiguchi T, Sugimoto H, Takehara K, Takagi H (June 1995). "Effect of biotin on ammonia intoxication in rats and mice". Journal of Gastroenterology. 30 (3): 351–5. doi:10.1007/bf02347511. PMID 7647902. S2CID 29888321.
- ^ Lichter-Konecki U, Nadkarni V, Moudgil A, Cook N, Poeschl J, Meyer MT, Dimmock D, Baumgart S (August 2013). "Feasibility of adjunct therapeutic hypothermia treatment for hyperammonemia and encephalopathy due to urea cycle disorders and organic acidemias". Molecular Genetics and Metabolism. 109 (4): 354–9. doi:10.1016/j.ymgme.2013.05.014. PMID 23791307.
- ^ Lichter-Konecki U, Caldovic L, Morizono H, Simpson K (April 2016). "Ornithine Transcarbamylase Deficiency". In Adam MP, Ardinger HH, Pagon RA, Wallace SE, Bean LJ, Stephens K, Amemiya A (eds.). GeneReviews. Seattle (WA): University of Washington, Seattle. PMID 24006547.
Further reading[edit]
- Tuchman M, Plante RJ (1995). "Mutations and polymorphisms in the human ornithine transcarbamylase gene: mutation update addendum". Human Mutation. 5 (4): 293–5. doi:10.1002/humu.1380050404. PMID 7627182. S2CID 2951786.
- Tuchman M (1993). "Mutations and polymorphisms in the human ornithine transcarbamylase gene". Human Mutation. 2 (3): 174–8. doi:10.1002/humu.1380020304. PMID 8364586. S2CID 42060839.
- Matsuda I, Tanase S (September 1997). "The ornithine transcarbamylase (OTC) gene: mutations in 50 Japanese families with OTC deficiency". American Journal of Medical Genetics. 71 (4): 378–83. doi:10.1002/(SICI)1096-8628(19970905)71:4<378::AID-AJMG2>3.0.CO;2-Q. PMID 9286441.
- Wakabayashi Y (July 1998). "Tissue-selective expression of enzymes of arginine synthesis". Current Opinion in Clinical Nutrition and Metabolic Care. 1 (4): 335–9. doi:10.1097/00075197-199807000-00004. PMID 10565370.
- Tuchman M, Jaleel N, Morizono H, Sheehy L, Lynch MG (February 2002). "Mutations and polymorphisms in the human ornithine transcarbamylase gene". Human Mutation. 19 (2): 93–107. doi:10.1002/humu.10035. PMID 11793468. S2CID 25848898.
- Feldmann D, Rozet JM, Pelet A, Hentzen D, Briand P, Hubert P, Largilliere C, Rabier D, Farriaux JP, Munnich A (July 1992). "Site specific screening for point mutations in ornithine transcarbamylase deficiency". Journal of Medical Genetics. 29 (7): 471–5. PMC 1016021. PMID 1353535.
- Tuchman M, Holzknecht RA, Gueron AB, Berry SA, Tsai MY (November 1992). "Six new mutations in the ornithine transcarbamylase gene detected by single-strand conformational polymorphism". Pediatric Research. 32 (5): 600–4. doi:10.1203/00006450-199211000-00024. PMID 1480464.
- Dawson SJ, White LA (May 1992). "Treatment of Haemophilus aphrophilus endocarditis with ciprofloxacin". The Journal of Infection. 24 (3): 317–20. doi:10.1016/S0163-4453(05)80037-4. PMID 1602151.
- Suess PJ, Tsai MY, Holzknecht RA, Horowitz M, Tuchman M (June 1992). "Screening for gene deletions and known mutations in 13 patients with ornithine transcarbamylase deficiency". Biochemical Medicine and Metabolic Biology. 47 (3): 250–9. doi:10.1016/0885-4505(92)90033-U. PMID 1627356.
- Grompe M, Caskey CT, Fenwick RG (February 1991). "Improved molecular diagnostics for ornithine transcarbamylase deficiency". American Journal of Human Genetics. 48 (2): 212–22. PMC 1683033. PMID 1671317.
- Hentzen D, Pelet A, Feldman D, Rabier D, Berthelot J, Munnich A (December 1991). "Fatal hyperammonemia resulting from a C-to-T mutation at a MspI site of the ornithine transcarbamylase gene". Human Genetics. 88 (2): 153–6. doi:10.1007/bf00206063. PMID 1721894. S2CID 23113285.
- Strautnieks S, Rutland P, Malcolm S (December 1991). "Arginine 109 to glutamine mutation in a girl with ornithine carbamoyl transferase deficiency". Journal of Medical Genetics. 28 (12): 871–4. doi:10.1136/jmg.28.12.871. PMC 1017166. PMID 1757964.
- Carstens RP, Fenton WA, Rosenberg LR (June 1991). "Identification of RNA splicing errors resulting in human ornithine transcarbamylase deficiency". American Journal of Human Genetics. 48 (6): 1105–14. doi:10.1016/j.devcel.2014.12.007. PMC 1683104. PMID 2035531.
- Hata A, Matsuura T, Setoyama C, Shimada K, Yokoi T, Akaboshi I, Matsuda I (May 1991). "A novel missense mutation in exon 8 of the ornithine transcarbamylase gene in two unrelated male patients with mild ornithine transcarbamylase deficiency". Human Genetics. 87 (1): 28–32. doi:10.1007/BF01213087. PMID 2037279. S2CID 31384734.
- Legius E, Baten E, Stul M, Marynen P, Cassiman JJ (August 1990). "Sporadic late onset ornithine transcarbamylase deficiency in a boy with somatic mosaicism for an intragenic deletion". Clinical Genetics. 38 (2): 155–9. doi:10.1111/j.1399-0004.1990.tb03565.x. PMID 2208768. S2CID 21521531.
- Finkelstein JE, Francomano CA, Brusilow SW, Traystman MD (June 1990). "Use of denaturing gradient gel electrophoresis for detection of mutation and prospective diagnosis in late onset ornithine transcarbamylase deficiency". Genomics. 7 (2): 167–72. doi:10.1016/0888-7543(90)90537-5. PMID 2347583.
- Grompe M, Muzny DM, Caskey CT (August 1989). "Scanning detection of mutations in human ornithine transcarbamoylase by chemical mismatch cleavage". Proceedings of the National Academy of Sciences of the United States of America. 86 (15): 5888–92. Bibcode:1989PNAS...86.5888G. doi:10.1073/pnas.86.15.5888. PMC 297736. PMID 2474822.
- Lee JT, Nussbaum RL (December 1989). "An arginine to glutamine mutation in residue 109 of human ornithine transcarbamylase completely abolishes enzymatic activity in Cos1 cells". The Journal of Clinical Investigation. 84 (6): 1762–6. doi:10.1172/JCI114360. PMC 304053. PMID 2556444.
- Chu TW, Eftime R, Sztul E, Strauss AW (June 1989). "Synthetic transit peptides inhibit import and processing of mitochondrial precursor proteins". The Journal of Biological Chemistry. 264 (16): 9552–8. doi:10.1016/S0021-9258(18)60567-8. PMID 2722850.
- Hata A, Setoyama C, Shimada K, Takeda E, Kuroda Y, Akaboshi I, Matsuda I (July 1989). "Ornithine transcarbamylase deficiency resulting from a C-to-T substitution in exon 5 of the ornithine transcarbamylase gene". American Journal of Human Genetics. 45 (1): 123–7. PMC 1683378. PMID 2741942.
- Summar ML, Tuchman M (29 April 2003). "Urea Cycle Disorders Overview". GeneReviews [Internet]. PMID 20301396.









